How it works
The vairous parameters are measured independently in the analysis pipeline and the user can select those analysis which are required for the given patient.
What is the array quality control?
Heber and Sick (Heber et al, 2006 - see literature) suggested eight quality metrics as a basic quality assessment for Affymetrix microarrays. First we have implemented their methods and tested them on an extended version of our previously published database (Gyorffy et al, 2010 - see literaure). The distribution of the arrays was assessed and outliers were identified as those having a parameter value outside of the range of 95% of samples. Then, the "Array quality control" parameter implemented in www.recurrenceonline.com was set to give a warning in cases in which the thresholds published by Heber et al. are surpassed or outliers are detected as compared to our meta-analysis.
What is the risk category using the stronges genes?
Using our validation database we identified the genes with the strongest predictive power in all patients, in lymph node positive, and in lymph node negative ER positive and ER negative patients. First, a filtering was performed to select only those probe sets which reliably work on the microarray. Probe sets were retained having a median expression over 890 (the whole-array median) or having a median expression of at least 445 and covering at least 20% of the gene and not mapping to multiple genes. After this, the gene with the lowest p value and the highest HR value in the given cohort of patients was selected. Then, the second probe set was added, and the mean expression of the two probe sets was used for classification. This was repeated as long as the predictive power of the mean of the used probe sets increased. A leave-one-out cross validation (LOOVC) was performed in each of the cohorts to measure the robustness as whether the same genes will be selected by excluding any of the samples.
What is the recurrence score?
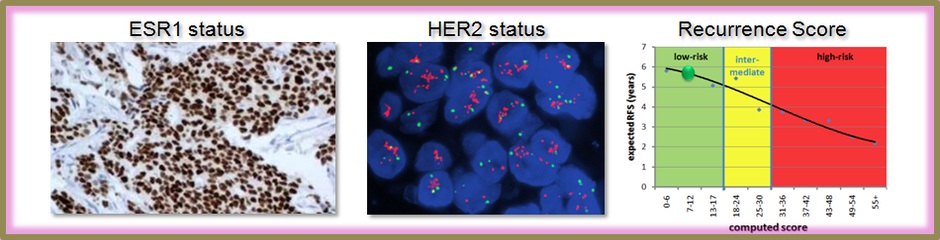
Recently, an RT-PCR based multigene assay investigating 21 genes was suggested to predict distant recurrence in patients with breast cancer who have no involved lymph nodes and estrogen-receptor positive tumors (Paik et al, 2004 - see Literature). In this, a gene-expression based recurrence score is computed which divides patients into groups having high/intermediate/low risk of recurrence. The assay provides treatment decision support as to which patient should receive adjuvant chemotherapy (Paik et al, 2006 - see Literature). A genome-wide Affymetrix microarray measures over 22000 genes including those 21 suggested to be used for breast cancer classification. We developed an online analysis tool to compute the recurrence score using gene expression data from Affymetrix microarrays.
How is the recurrence score computed?
After an initial quality control the raw Affymetrix .CEL files are MAS5 normalized. The differences of the log-transformed expression of the 16 genes and the housekeeping genes ACTB, GAPDH, RPLP0, GUS and TFRC are subtracted from the range top (adjustable parameter) to emulate RT-PCR results. For genes with multiple probe sets available on the Affymetrix microarrays the average expression, the probe set with the highest average expression can be used (adjustable parameter). Then, the recurrence score is computed as described by Paik et al, 2004. Finally, samples are classified as belonging to the high/intermediate/low group based on their recurrence score.
How is the ER status computed?
Gong et al demonstrated the feasibility to measure estrogen receptor and ERBB2 status reliably and reproducibly using Affymetrix microarrays (Gong et al, 2007 - see Literature). We have implemented their approach using the suggested cutoff values of 500 (in the probe set 205225_at) for estrogen receptor.
How is the HER2 status computed?
For the ERBB2 receptor the bimodal distribution of the validation datasets was decomposed into two Gaussian distributions, which correspond to two specific ERBB2 expression statuses. Based on the two inferred distributions a cohort-specific cut-off value for ERBB2 using Mahalanobis distance - which minimizes the estimated false positive rate (FPR) and the false negative rate (FNR) - was derived (Li et al, 2010 - see literature). The actual cutoff for ERBB2 is user selectable: "bimodal distribution" uses 4,800 as cutoff, while "immunhistochemistry" uses the 1,150 cutoff suggested by Gong et al.
CITED LITERATURE
Gong Y, Yan K, Lin F, et al.: Determination of oestrogen-receptor status and ERBB2 status of breast carcinoma: a gene-expression profiling study. Lancet Oncol. 2007 Mar;8(3):203-11. Lancet link.
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z.: An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010 Oct;123(3):725-31. Springer link.
Gyorffy and Schafer: Meta-analysis of gene expression profiles related to relapse-free survival in 1,079 breast cancer patients. Breast Cancer Res Treat. 2009 Dec;118(3):433-41. Springer link.
Gyorffy B, Molnar B, Lage H, Szallasi Z, Eklund AC.: Evaluation of microarray preprocessing algorithms based on concordance with RT-PCR in clinical samples. PLoS One. 2009 May 21;4(5):e5645. Free full text.
Heber S, Sick B. Quality assessment of Affymetrix GeneChip data. OMICS 2006;10:358-68.
Li Q, Eklund AC, Juul N, Haibe-Kains B, Workman CT, Richardson AL, Szallasi Z, Swanton C. Minimising Immunohistochemical False Negative ER Classification Using a Complementary 23 Gene Expression Signature of ER Status. PLoS One. 2010 Dec 1;5(12):e15031. Free full text.
Paik S, Shak S, Tang G, et al: A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351:2817-2826, 2004. Free full text.
Paik S, Tang G, Shak S, et al.: Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J Clin Oncol. 2006 Aug 10;24(23):3726-34. Full text.


